Single Molecule Studies of DNA-protein interactions
Mario Feingold
We use Optical Tweezers to manipulated single DNA molecules. This method can be used to probe various processes in which the DNA plays a role. In particular, we use this approach to study the interaction between the DNA and sequence specific proteins. Such protein will first search for the appropriate sequence on the DNA and once it has found it will initiate a binding process. These processes will influence the DNA that, in turn, will affect the position of the bead in the optical trap. The displacements that need to be observed in order to monitor such process are in the nanometer range and are the limit of the resolution of such a setup. On the other hand, one could amplify the effect by using a DNA that has several copies of the binding sequence.
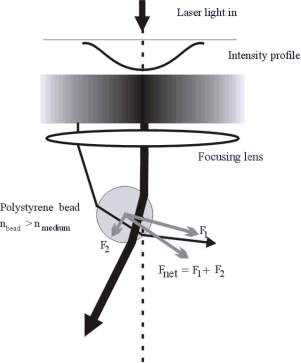
Figure: Optical Tweezers trap microbeads that can be attached to single DNA molecules. It is like having a handle at the end of the DNA.
Single Cell Dynamics
Mario Feingold
We use single cell phase-contrast and fluorescence time-lapse microscopy to monitor the morphological changes during the division of E. coli. To bypass the limitations of optical resolution, we process the images using pixel intensity values for edge detection. We study the dynamics of the constriction width, W, and find that its formation starts shortly after birth much earlier than can be detected by simply viewing phase-contrast images. A simple geometrical model is shown to reproduce the behavior of W(t). Moreover, the time-dependence of the cell length, L(t), consists of three linear regimes. The growth rates in the different regimes are related to each other and to the parameters of our model.
 |
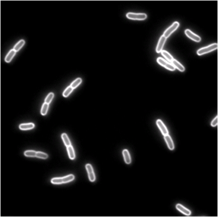 |
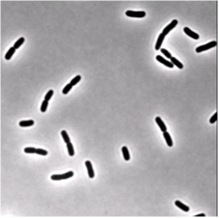
| Phase-contrast (right) and fluorescence (left- stained membrane) images of bacteria (E.coli). | |
 |
| This is life: 1.5 min (a), 8.6 min (b) and 19.4 min © from birth. In © the bacterium has just divided. |
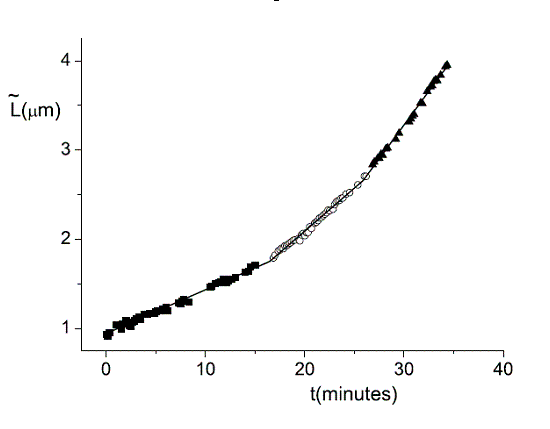 |
| During the lifetime of a bacterium it elongates linearly in three regimes. The transition between these regimes is rather sharp. Click to enlarge. |